TargetMol中国品牌商
12 年
手机商铺
- NaN
- 0
- 0
- 2
- 2
推荐产品
公司新闻/正文
TargetMol's 7 libraries for your research to combat against covid-19
771 人阅读发布时间:2020-03-20 16:22
TargetMol's 7 libraries for your research to combat against covid-19
| Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) belongs to a new evolutionary branch within the Coronaviruses (CoVs). The new emerging SARS-CoV-2 shares about 80% of the gene sequence of SARS-CoV; Proteins from SARS-CoV-2 and SARS-CoV were treated as homologous: identity value ≥ 65%. Therapeutics currently targeting spike RBD-ACE2, Nsp16, 3CLpro, PLpro, RdRp and X domain are possible treatments for SARS-CoV-2. |
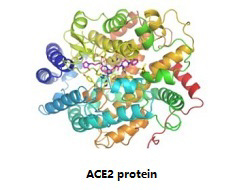 |
The prefusion SARS-CoV S1 subunit is structurally organized into four distinct domains: NTD, CTD1, CTD2 and CTD3. Among these, CTD1 is the receptor-binding domain, and one CTD1 in the trimer adopts an “up” conformation as a prerequisite for the binding of SARS-CoV to the cellular receptor angiotensin-converting enzyme 2 (ACE2). Similar observations of a protruding “up” CTD1 have also been reported for MERS-CoV S glycoproteins. Targeting the interaction of S protein RBD and ACE2 is an important therapeutic strategy to block coronavirus from entering host cells. |
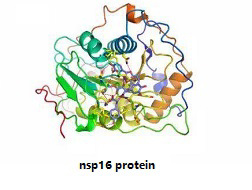 |
Nsp16 provides the viral mRNA with the ability to camouflage and obscure itself from the host cell by catalyzing methylation on m7GpppA-RNA, thus preventing recognition and activation of the host immune response which is essential for successful viral infection. Nsp16/nsp10 interface may represent a better drug target than the viral MTase active site for developing highly specific anti-coronavirus drugs. New evidence from a joint research team showed that Saquinavir and Lopinavir, two anti-HIV drugs, can inhibit the activity of NSP16 methyltransferase, thereby suppressing the function and replication of the virus. |
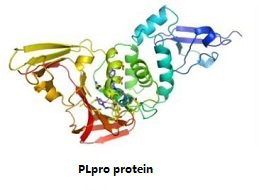 |
Papain-like proteinase (PLpro), encoded in nsp3, is responsible for the cleavages of N-terminus of the replicase polyprotein to release Nsp1, Nsp2 and Nsp3, which is essential for correcting virus replication and antagonizing the host’s innate immunity. As an indispensable enzyme in the process of coronavirus replication and infection of the host, PLpro has been a popular target for coronavirus inhibitors. |
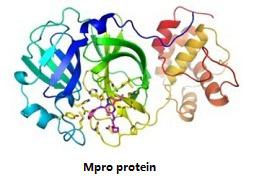 |
3CLpro, also known as Mpro or Nsp5, is a chymotrypsin-like cysteine protease that can proteolytically process the replicase polyprotein 1a and 1ab at their C-terminal regions to release Nsp4–Nsp16. Based on the protein structure of 3CLpro, multiple research papers published identified several drugs having the potential to treat COVID-19 by virtual screening, such as drugs that target PLpro and 3CLpro in other viruses such as HIV drugs, lopinavir and ritonavir. |
 |
RdRp (Nsp12) is an RNA-dependent RNA polymerase (RdRp). Except for reverse-transcribing viruses, all plus-strand RNA viruses encode an RdRp that functions as the catalytic subunit of the viral replication/transcription complex, directing viral RNA synthesis in concert with other viral proteins and, sometimes, host proteins. RdRp is the core component of virus genome replication system, and has been used as a very important drug target in the research of SARS-CoV and MERS-CoV inhibitors. Remdesivir, an effective anti-COVID-19, is a nucleotide analog inhibitor of RdRp. |
 |
X domain is a conserved structure of pp1a and becomes a part of nsp3 after pp1a cleaved by a virally encoded cysteine protease, the papain-like protease (PLpro). It is a catalytically active ADP-ribose-1″-phosphatase thought to play a role during synthesis of viral subgenomic RNAs thus can be used for drug screening or design. |



